| Line 15: | Line 15: | ||
<br><br> | <br><br> | ||
<br><br> | <br><br> | ||
| − | To import and export layouts for OptFlux check the plugins: | + | To import and export layouts for OptFlux check the plugins:<br> |
[[OptFlux3:CDG|CellDesigner Plugin]]<br> | [[OptFlux3:CDG|CellDesigner Plugin]]<br> | ||
[[OptFlux3:CYT|XGMML Plugin]]<br> | [[OptFlux3:CYT|XGMML Plugin]]<br> | ||
<br> | <br> | ||
Latest revision as of 12:50, 21 January 2013
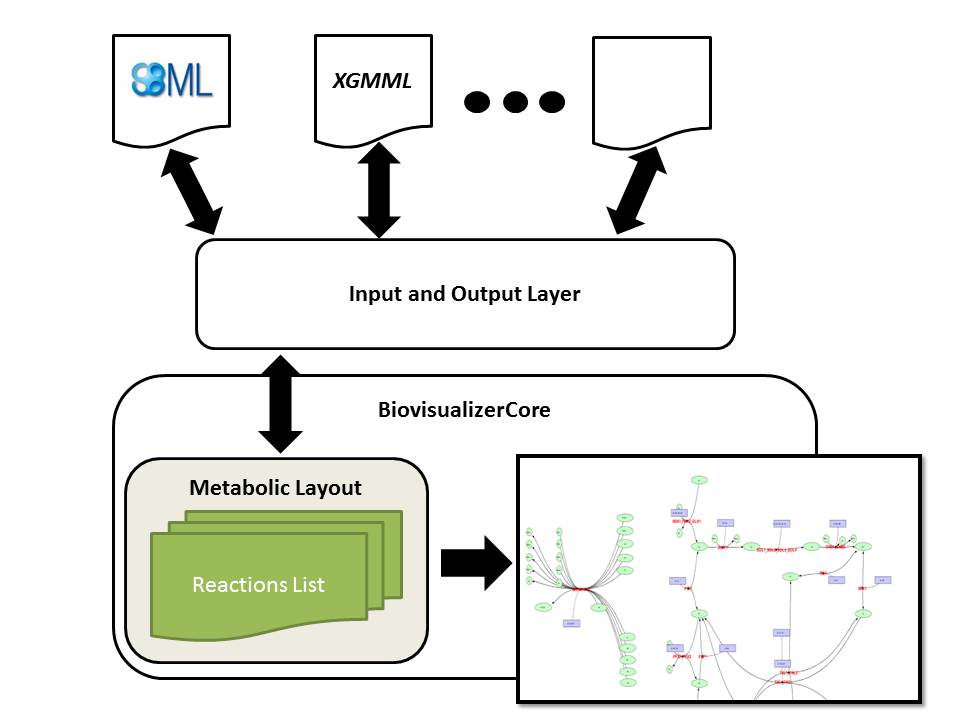
Biovisualizer Core provides a visual representation of a metabolic model. The network is represented as a graph, where biological entities (reactions and metabolites) are represented as nodes, and their interactions (consumption and production) as edges. The architecture of the visualizer library has two distinct layers: the BiovisualizerCore contains the specification of a visual representation of the metabolic network (layout), the tools to represent it in a 2D interface and methods for visual manipulation of the network; the second layer, the Input/Output layer, consists in the readers and writers for a variety of formats that can provide layouts to the BiovisualizerCore, working as a transformer receiving a file in a specific format, reading it and generating a layout.
The visualizer provides:
Visual filters Metabolic information Simulation results visual representation Elementary Flux Modes representation
To import and export layouts for OptFlux check the plugins:
CellDesigner Plugin
XGMML Plugin